Pedigree and fitness data for North American red squirrels (Tamiasciurus hudsonicus) from the Kluane Red Squirrel Project, a long-term field study conducted in the boreal forests of Yukon, Canada, running continuously since 1987. The project has individually marked and monitored thousands of squirrels across multiple generations, producing one of the most detailed wild-animal pedigrees available for quantitative genetic research.
red_squirrelsFormat
A data frame with 7,799 rows and 16 columns:
- personID
Integer. Unique individual identifier.
- momID
Integer. Mother's
personID.NAfor founders or individuals whose mother was not identified.- dadID
Integer. Father's
personID.NAfor founders or individuals whose father was not identified.- sex
Character. Sex:
"F"(female),"M"(male).- famID
Integer. Family group identifier assigned by
ped2fam.- byear
Integer. Birth year.
NAif unknown.- dyear
Integer. Death year.
NAif the individual's fate was not recorded or it was still alive at the end of the study.- lrs
Numeric. Lifetime reproductive success: total number of offspring surviving to independence across the individual's entire life.
NAif not available.- ars_mean
Numeric. Mean annual reproductive success across all years in which the individual was observed.
NAif no annual records are available.- ars_max
Numeric. Maximum ARS value recorded across all observed years.
- ars_med
Numeric. Median ARS value across all observed years.
- ars_min
Numeric. Minimum ARS value across all observed years.
- ars_sd
Numeric. Standard deviation of ARS values across all observed years.
- ars_n
Integer. Number of years for which an ARS value was recorded. Zero if the individual has no ARS records.
- year_first
Integer. First calendar year in which the individual was observed.
- year_last
Integer. Last calendar year in which the individual was observed.
Source
McFarlane, S.E., Boutin, S., Humphries, M.M., et al. (2015). Very low levels of direct additive genetic variance in fitness and fitness components in a red squirrel population. Dryad. doi:10.5061/dryad.n5q05
Details
Red squirrels in this population occupy individual year-round territories centered on a food cache (midden). Key fitness traits include annual reproductive success (ARS: the number of offspring surviving to independence in a given year) and lifetime reproductive success (LRS: the total number of such offspring over an individual's lifetime). These traits have been used to study the heritability of fitness in a natural population, with the original publication finding very low levels of direct additive genetic variance.
Family group IDs were assigned using ped2fam from
the BGmisc package. The original data are published under a
CC0 1.0 Universal Public Domain Dedication.
Examples
head(red_squirrels)
#> # A tibble: 6 × 16
#> personID momID dadID sex famID byear dyear lrs ars_mean ars_max ars_med
#> <dbl> <int> <int> <chr> <dbl> <int> <dbl> <int> <dbl> <dbl> <dbl>
#> 1 1 NA NA F 1 NA NA NA NA NA NA
#> 2 2 NA NA F 2 NA NA NA NA NA NA
#> 3 3 NA NA F 3 NA NA NA NA NA NA
#> 4 4 NA NA F 4 NA NA NA NA NA NA
#> 5 5 NA NA F 5 NA NA NA NA NA NA
#> 6 6 NA NA F 6 NA NA NA NA NA NA
#> # ℹ 5 more variables: ars_min <dbl>, ars_sd <dbl>, ars_n <dbl>,
#> # year_first <dbl>, year_last <dbl>
str(red_squirrels)
#> tibble [7,799 × 16] (S3: tbl_df/tbl/data.frame)
#> $ personID : num [1:7799] 1 2 3 4 5 6 7 8 9 10 ...
#> $ momID : int [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ dadID : int [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ sex : chr [1:7799] "F" "F" "F" "F" ...
#> $ famID : num [1:7799] 1 2 3 4 5 6 7 8 9 10 ...
#> $ byear : int [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ dyear : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ lrs : int [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_mean : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_max : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_med : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_min : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_sd : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ ars_n : num [1:7799] 0 0 0 0 0 0 0 0 0 0 ...
#> $ year_first: num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
#> $ year_last : num [1:7799] NA NA NA NA NA NA NA NA NA NA ...
# LRS distribution by sex
tapply(red_squirrels$lrs, red_squirrels$sex, summary)
#> [[1]]
#> Min. 1st Qu. Median Mean 3rd Qu. Max. NAs
#> NA NA NA NaN NA NA 1
#>
#> $`0`
#> Min. 1st Qu. Median Mean 3rd Qu. Max. NAs
#> NA NA NA NaN NA NA 1
#>
#> $F
#> Min. 1st Qu. Median Mean 3rd Qu. Max. NAs
#> 0.000 0.000 0.000 1.406 0.000 31.000 1552
#>
#> $M
#> Min. 1st Qu. Median Mean 3rd Qu. Max. NAs
#> 0.0000 0.0000 0.0000 0.3361 0.0000 17.0000 3262
#>
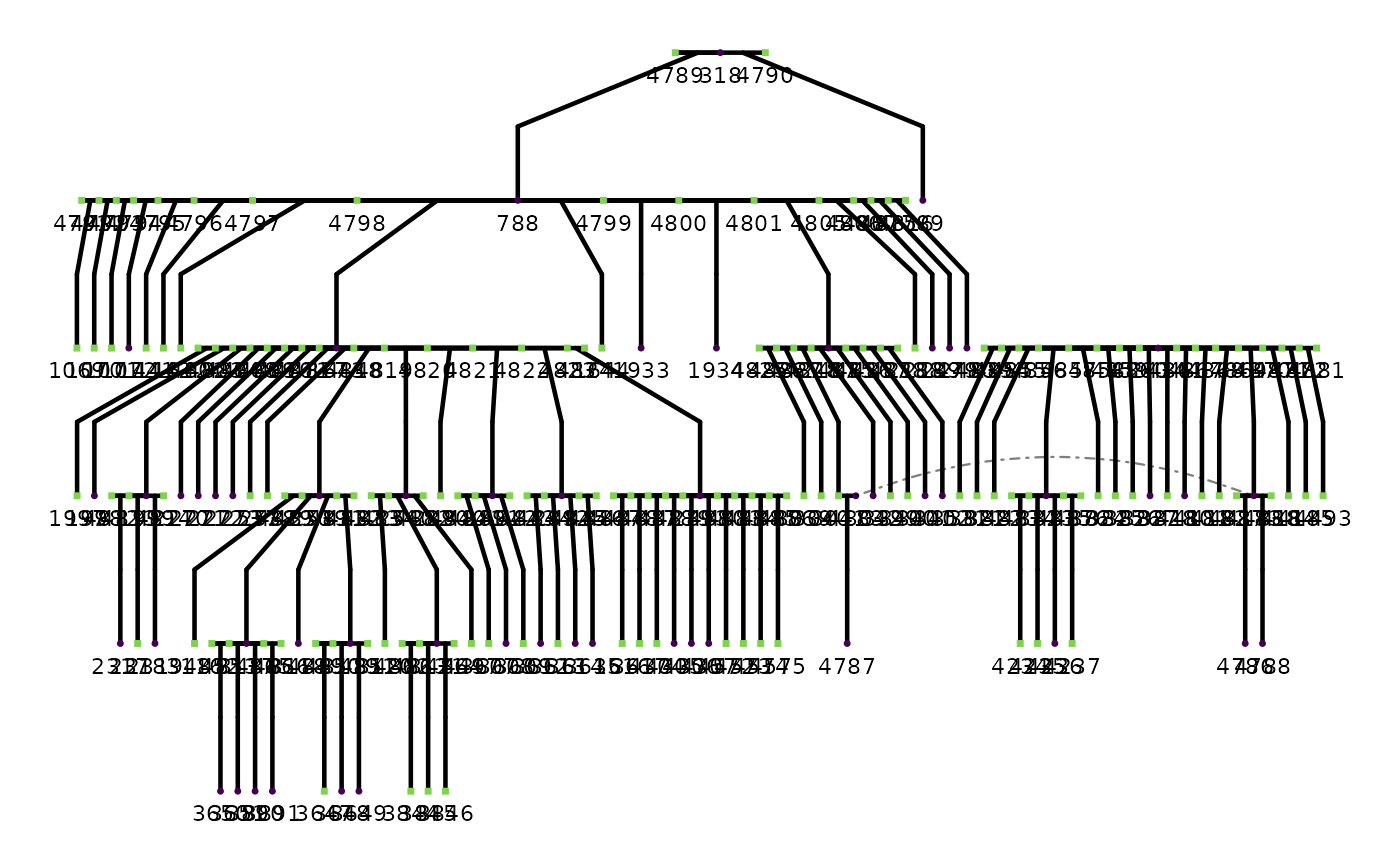
# Single-family pedigree plot
if (requireNamespace("ggpedigree", quietly = TRUE)) {
fam160 <- subset(red_squirrels, famID == 160)
ggpedigree::ggPedigree(fam160,
personID = "personID", momID = "momID", dadID = "dadID",
config = list(add_phantoms = TRUE, code_male = "M")
)
}
#> REPAIR IN EARLY ALPHA